Organismal fitness if often constrained by interactions with other species, such as parasites, pathogens or mutualists. To understand the evolutionary impact of such interactions, we analysed host-pathogen as well as putative beneficial interactions between insects and bacteria.
Insect–gut bacterial associations
Apart from detrimental host-pathogen interactions, a large body of work shows that insects have diverse beneficial associations with their gut microbial communities. However, these interactions are relatively poorly understood in natural insect populations, and key predictions regarding their evolution and impact remain unanswered. For instance, an evolved host-microbiome relationship should generally result in a temporally and spatially stable microbial community, with host-imposed filtering mechanisms leading to non-neutral community assembly. However, we found that gut bacterial communities of wild-caught dragonflies show large variation across space, season and host taxonomy (Nair and Agashe 2016), as well as strong signatures of neutral community assembly (Deb et al 2019). Interestingly, gut content analysis revealed that this variability likely arises due to dietary specialization across host species, such that diet-acquired microbes largely dictate community assembly, with little host-imposed filtering.
Similarly, in wild-caught butterflies, we find that host plant-derived microbes dominate the microbiomes of larval stages, and there is surprisingly little change in this community during the large dietary shifts that occur during development (phytophagous larvae develop into nectar- or plant sap-feeding adults; Phalnikar et al 2018). Together, our work suggests that neither dragonflies nor butterflies have evolved significant and stable associations with their gut microbiomes. We tested this hypothesis by experimentally manipulating larval gut microbiomes; and found very weak fitness effects of depleting larval microbiomes in butterflies (Phalnikar et al 2019).
Flour beetles are an interesting contrast to the weak host-microbiome associations we found in dragonflies and butterflies. In this case, laboratory rearing conditions closely mimic the environment to which beetles have adapted for the past thousands of years: stored grain warehouses. We found that eliminating microbes from wheat flour reduces the fitness of wild-collected wheat-adapted beetles, whereas microbes do not affect beetle fitness in novel flours. However, as these beetles adapted to new flours in the laboratory for 2 years (~10 generations), they evolved a dependence on flour-associated bacteria, with a concomitant change in their gut microbiome (Agarwal and Agashe, in review). This is potentially the first experimental demonstration of an evolved host-microbiome association, and is remarkable for the very short time in which it occurred. Our work paves the way for addressing major open questions about host-gut microbiome interactions: their mechanistic basis, and the joint evolutionary impacts on the host, the bacterial community, and bacterial genomes.
The evolution of rice-associated Methylobacterium
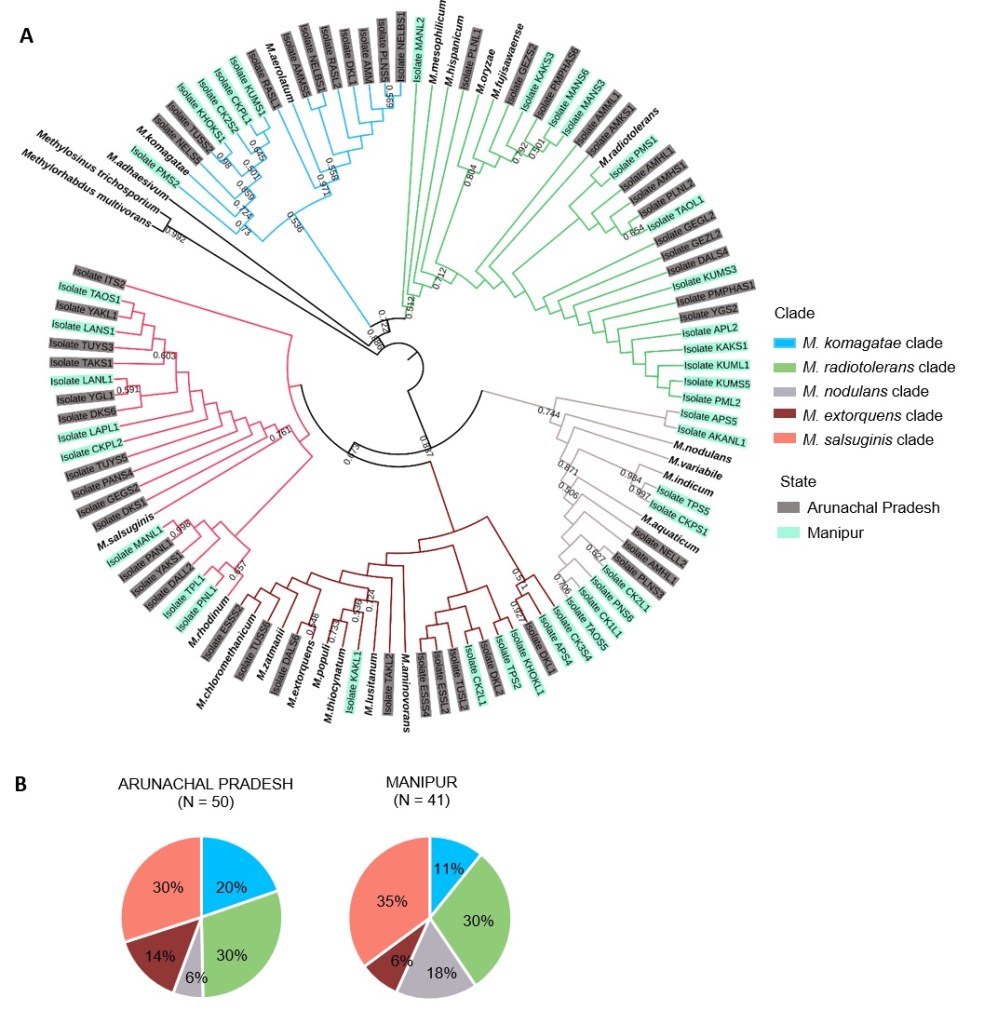
We are testing the impact of host association on Methylobacterium species isolated from the leaves of traditionally cultivated rice landraces from North-Eastern India. Previously published work suggests that some species from this genus (e.g. M. oryzae and M. phyllospherae) use methanol emitted by rice leaves as a primary carbon source, and may produce growth hormones that enhance host growth. We found that phenotypic diversity in Methylobacterium is associated strongly with the host landrace identity, rather than other factors (Sanjenbam et al 2020). We are now testing whether the genetically distinct rice varieties in northeastern India have co-evolved specific associations with their local Methylobacterium partners. If we find specific associations between local rice and associated Methylobacterium species, we aim to characterize the chemical and genetic basis of the association. This work is conducted in collaboration with P. V. Shivaprasad and Radhika Venkatesan at NCBS.
